The Chromium Controller from 10X Genomics is designed to automate parallel sample partitioning and molecular barcoding at the single cell level. The controller allows for single cell analysis of CNV, gene expression, immune profiling, and ATAC.
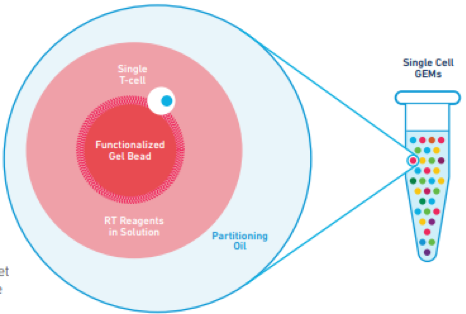 Gel Beads, each containing millions of barcoded oligonucleotides, are mixed with cells or nuclei from a sample pool at a flow rate designed to enable each Gel Bead to bind to a single cell or nucleus. Once bound, the Gel Beads are immersed in an oil solution, capturing each single bead, cell, and the immediately surrounding reagents in a bubble which then serves as a reaction vesicle, shown right.
Gel Beads, each containing millions of barcoded oligonucleotides, are mixed with cells or nuclei from a sample pool at a flow rate designed to enable each Gel Bead to bind to a single cell or nucleus. Once bound, the Gel Beads are immersed in an oil solution, capturing each single bead, cell, and the immediately surrounding reagents in a bubble which then serves as a reaction vesicle, shown right.
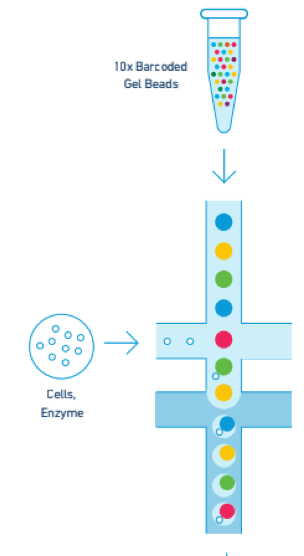 In the oil bubble, known as a GEM (Gel bead in EMulsion), the bead is dissolved and RNA is extracted from the cell or nucleus, reverse transcribed, and barcoded with the unique barcode sequences found inside each Gel Bead.
In the oil bubble, known as a GEM (Gel bead in EMulsion), the bead is dissolved and RNA is extracted from the cell or nucleus, reverse transcribed, and barcoded with the unique barcode sequences found inside each Gel Bead.
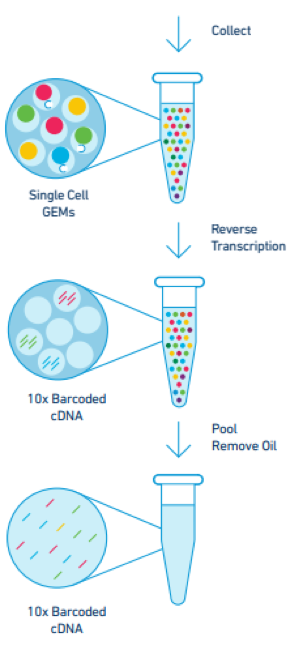 The cDNA from all the cells in the original sample are then pooled and the oil is removed, leaving behind barcoded cDNA that can be given Illumina adapters for sequencing. Following sequencing, the unique barcode tags from the Gel Beads enable the transcripts from each individual cell or nucleus to be grouped together during analysis.
The cDNA from all the cells in the original sample are then pooled and the oil is removed, leaving behind barcoded cDNA that can be given Illumina adapters for sequencing. Following sequencing, the unique barcode tags from the Gel Beads enable the transcripts from each individual cell or nucleus to be grouped together during analysis.
Please contact the Genomics Facility in advance if you wish to run samples on the 10X platform and ensure that your cells are prepared to the recommended specifications before the run, as the reagents used are time-sensitive. Also, note that while cells are barcoded individually, up to eight sample pools can be run in parallel.
- Single-Cell Next Generation Sequencing
Please see our rates page for detailed pricing information.



