The MiSeq used to sequence NGS and microbiome libraries. The run modules available are a nano-kit with 1 million reads (2x150 or 2x250), version 2 with 12-15 million reads (2x150 and 2x250) and version 3 with 22-25 million reads (2x75 and 2x300).
The NextSeq instrument from Illumina uses sequencing-by-synthesis (SBS) chemistry to accurately determine the sequences of millions of unique DNA fragments at a time. During each sequencing cycle, reversibly-labeled nucleotides are washed across a surface of oligo clusters, allowing one nucleotide to bind to the growing complement to each individual oligo strand. The labeled nucleotides are imaged, and the dye is then chemically cleaved to allow the next nucleotide in the sequence to bind. Base calls are determined using the signal intensity measurements taken in the imaging step of each cycle.
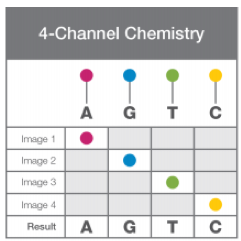 While all Illumina instruments use SBS chemistry, there are multiple dye combinations used to capture the bases in each cycle. On the MiSeq, a 4-channel chemistry is used. As shown on the right, each base is tagged with a unique dye which fluoresce in response to a unique wavelength; after the clusters have been washed with the tagged bases, therefore, the flowcell is imaged four times per cycle to identify the clusters in which each base has been incorporated.
While all Illumina instruments use SBS chemistry, there are multiple dye combinations used to capture the bases in each cycle. On the MiSeq, a 4-channel chemistry is used. As shown on the right, each base is tagged with a unique dye which fluoresce in response to a unique wavelength; after the clusters have been washed with the tagged bases, therefore, the flowcell is imaged four times per cycle to identify the clusters in which each base has been incorporated.
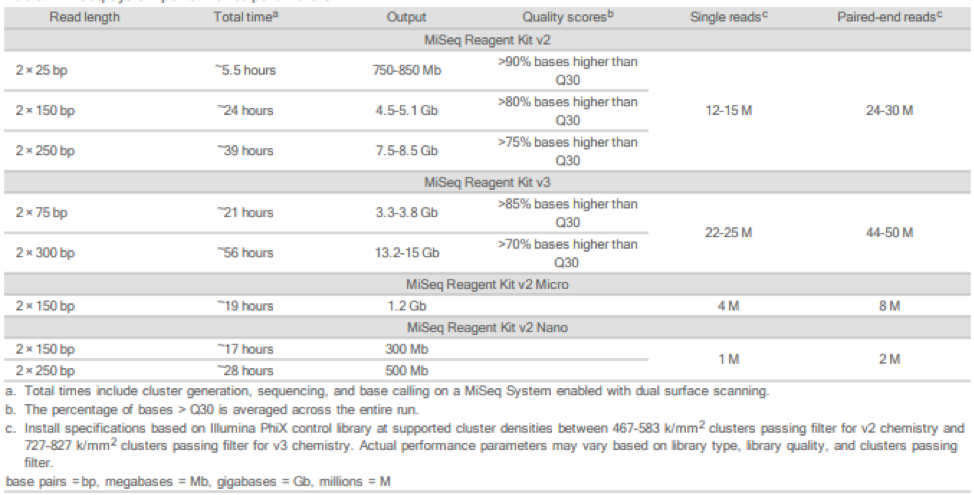
The different run modules for the MiSeq, with their parameters and expected quality, are outlined in the following table from Illumina. Please note that the Run Time refers to on-instrument time and not turn-around time for run processing at the facility (which includes sample QC and data conversion as well, and is dependent on the presence of other runs in the queue).
At the Genomics Facility, we have found that the MiSeq is the ideal platform for microbiome sequencing and targeted resequencing. Past projects include metagenomic sequencing with Biodesign’s Swette Center for Environmental Biotechnology and genetic analysis of ancient and modern human populations via targeted resequencing with the Stone Lab at ASU
- Next generation sequencing
Please see our rates page for detailed pricing information.






